Controlling intracellular signal network dynamics with light
Optogenetic investigations by the Dehmelt group reveal mechanism for regulation of cell contraction dynamics
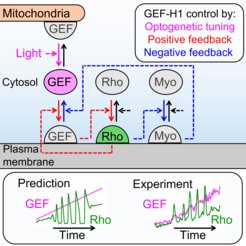
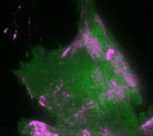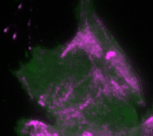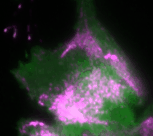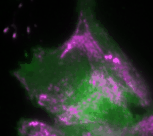
In the new study, the Dehmelt group uses microscopy-based, light-switchable perturbations to specifically and acutely manipulate a signal network that controls cell contractions in individual, living cells. At the same time, the dynamic response of this signal network was measured to investigate the mechanism, how these perturbations are processed by cells. Based on these results, a theoretical model was derived which quantitatively describes the components and interactions of this signal network. A central aspect of this system are positive and negative feedback loops, which generate highly dynamic pulses of the cell contraction regulator Rho and the molecular motor myosin (Video 1). The theoretical model of the Dehmelt group made it possible to predict how the dynamics of this signal network would react to specific perturbations of these feedback loops. These predictions were confirmed by controlling a key signal network component with light in individual, living cells. Figure 1 illustrates causal connections in this signal network and how light was used to manipulate the concentration of one key component, the Rho activator GEF-H1, via “optogenetic tuning”. With this technique it was discovered that this signal network generates particularly intense pulses of the cell contraction regulator Rho at intermediate concentrations of GEF-H1. In contrast, the sensitivity of the signal network to process physiologically relevant mechanical and biochemical signals is maximal at low concentrations of this network component. These new theoretical and experimental findings form the basis for a better understanding of more complex processes in embryonic development and tumor progression.

