
Sabrina Pospich
Project Group Leader, Structural Biochemistry
Mechanistic Structural Biology
News
Are you interested in conducting science at the interface of biochemistry and physics? We have an open master position. With your help, we want to elucidate the 3D structure of silk and better understand the spinning process. Your tasks would include rearing caterpillars, extracting silk, systematically breaking silk down into its components and analysing them.
Research focus
When talking about silk most people directly think of a luxurious piece of cloth. However, silk is much more than just a delicate fabric. Silks consist of proteins, primarily of so-called spidroins and fibroins, which are assembled into silk fibers in a complex spinning process in specialized ducts. The silks produced by spiders and silkworms have remarkable mechanical properties competing with those of steel and Kevlar. Additionally, they are biocompatible and biodegradable making them a super-strong light-weight green material. The enormous potential of silks and silk-inspired synthetic materials for modern biomedicine and material science has been recognized early, resulting in various patents and companies offering synthetic silk-like fibers. However, there is surprisingly scarce data on the structure-function relationship of silk proteins as well as the spinning process itself. This is mostly due to the large size (~200-350 kDa), the highly repetitive amino acid sequence and the unusual semi-crystalline structure of silk proteins, which impede the characterization by traditional methods such as X-ray crystallography and in solution nuclear-magnetic resonance spectroscopy.
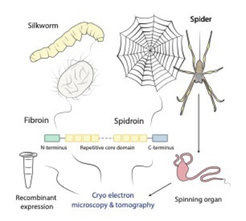
The aim of our group is to study the three-dimensional structure of silks and the associated spinning process by latest high-resolution cryo electron microscopy and tomography techniques. For this purpose, we pursue two complementary approaches. On the one hand, we study different silk types produced by live silkworms and spiders as well as their spinning ducts. On the other hand, we recombinantly express synthetic spidroins to characterize the role of specific domains and the proteins’ structural transition during artificial spinning.
Knowledge gained from this research project has the potential to promote the development of silk-based, ecological high-tech fibers, which could meet the urgent need for sustainable materials.
Selected publications
Oosterheert, W., Klink, B. U., Belyy, A., Pospich, S., and Raunser, S. (2022) Structural basis of actin filament assembly and aging. Nature, 611(7935):374-379
Wang, Z.*, Grange, M.*, Pospich, S., Wagner, T., Kho, A. L., Gautel, M. and Raunser, S. (2022) Structures from intact myofibrils reveal mechanism of thin filament regulation through nebulin. Science, 375(6852): eabn1934.
Pospich, S., Sweeney, H.L. , Houdusse, A. and Raunser, S. (2021). High-resolution structures of the actomyosin-V complex in three nucleotide states provide insights into the force generation mechanism. eLife 10:e73724; doi: 10.7554/eLife.73724.
Pospich, S., Küllmer, F., Nasufovic, V., Funk, J., Belyy, A., Bieling, P., Arndt, H.-D. and
Raunser, S. (2021). Cryo-EM resolves molecular recognition of an optojasp photoswitch bound to actin filaments in both switch states. Angewandte Chemie (International ed. In English), 60(16):8678-8682.
Pospich, S., Merino, F. and Raunser, S. (2020). Structural Effects and Functional Implications of Phalloidin and Jasplakinolide Binding to Actin Filaments. Structure, 28(4):437–449.e5.
Merino, F., Pospich, S. and Raunser, S. (2019). Towards a structural understanding of the remodeling of the actin cytoskeleton. Seminars in cell & developmental biology, 102:51–64.
Pospich, S. and Raunser, S. (2018). Single particle cryo-EM-an optimal tool to study cytoskeletal proteins. Current opinion in structural biology, 52:16–24.
Merino, F.*, Pospich, S.*, Funk, J., Wagner, T., Küllmer, F., Arndt, H.-D., Bieling, P. and Raunser, S. (2018). Structural transitions of F-actin upon ATP hydrolysis at near-atomic resolution revealed by cryo-EM. Nature structural & molecular biology, 25(6):528–537.
Pospich, S., Kumpula, E.-P., von der Ecken, J., Vahokoski, J., Kursula, I. and Raunser, S. (2017). Near-atomic structure of jasplakinolide-stabilized malaria parasite F-actin reveals the structural basis of filament instability. Proceedings of the National Academy of Sciences of the United States of America, 114(40):10636–10641.
