Structural Biochemistry
Identifying the 3D structure of proteins and their interactions with other molecules is crucial for understanding their function and underlying mechanisms. Proteins are highly complex molecules that perform a wide range of functions in cells and play many critical roles. By unravelling their structures at a molecular level, we are able to take steps towards better understanding the complex processes involved, ultimately helping to map out a complete picture of their function and regulation in cells in health and disease.
In our department, we utilize powerful techniques for studying the structures of proteins, protein complexes and cellular compartments: electron cryomicroscopy (cryo-EM) and electron cryotomography (cryo-ET). By freezing samples in a hydrated state and imaging them with a high-end electron microscope, we obtain high-resolution images that reveal the molecular details of cells and the proteins they contain at an atomic level. We are then able to use these images to build a comprehensive 3D model of single proteins or complex cellular subregions, which provide invaluable information on how proteins function and interact with their environment.
Along a list of milestones and remarkable discoveries thanks to the continuous dedication and hard work of our students, staff and scientists, a few highlights of our achievements include: the determination of cryo-EM structures of essential muscle and cytoskeletal proteins, including F-actin, the actin-tropomyosin complex and the cytoplasmic actin-tropomyosin-myosin complex at unprecedented resolution. We applied cryo-ET in order to reveal the detailed three-dimensional architecture of the sarcomere, the basic contractile unit of skeletal and heart muscle cells. In another recent achievement, we determined for the first time a structure of nebulin in intact sarcomeres at 4.5 Å. This structure has profound implications in unravelling the mechanisms involved in muscle structure and function in health and disease, since it reveals nebulin as a “molecular ruler” and stabiliser of the thin filament.
Zooming in on Muscle Cells
Watch how an international Team led by Prof. Stefan Raunser has produced the first high-resolution 3D image of the sarcomere, the basic contractile unit of skeletal and heart muscle cells, by using electron cryo-tomography (cryo-ET).
Our primary focus in the upcoming years will be on determining the high-resolution structure of sarcomeres and their components from different organisms. A precise molecular understanding of how the entire sarcomere machine forms and functions will allow us to better understand its role in health, disease and ageing.
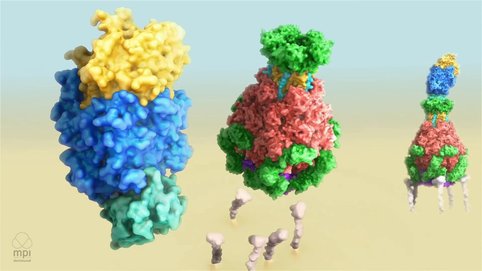
One of the major discoveries of our department involves understanding the hidden mechanism of action of ABC-type toxins. In particular, we revealed that Tcs (tripartite ABC-type bacterial toxin complexes from Photorhabdus luminescens) use a special syringe-like device for cell entry and we deciphered the molecular mechanism of intoxication involving toxin activation, receptor binding, membrane permeation and protein translocation. The hallmark of our work was the demonstration that Tc toxins can in fact turn into molecular syringes for delivering proteins of interest across membranes. We are currently working towards the molecular engineering of such toxins in order to transform them into potential medical nanosyringes.
Mechanism of Tc Toxin Action
State-of-the-art hardware and software development is the backbone of our research, offering the necessary support for our biological work. To this end, we successfully developed the SPHIRE program which offers users an easy-to-use interface for non-standard protein complexes such as helical filaments or flexible protein complexes, which are difficult to work with. The latest package of SPHIRE features a fully automated procedure for single particle analysis with quality control checkpoints, helical SPHIRE and per-particle CTF refinement called tranSPHIRE. We also developed crYOLO, which is based on deep learning algorithms and enables the automatic picking of particles. crYOLO is one of the most widely-used computer programs for particle selection world-wide. Building on this success, our department has further developed TomoTwin: a robust, first-in class general picking model for cryo-electron tomograms based on deep metric learning. TomoTwin is the first program that allows users to identify proteins in tomograms de novo without manually creating training data, setting new standards in cryo-ET data processing. In the next years, we aim to develop novel technology and software beyond the current state-of-the-art in terms of in situ structural cell biology in order to establish a modern approach that allows the study of cardiac disease at the molecular level using cryo-ET.
> More information on the research of the Raunser Group